Genome Sequencing Study Finds Clues to Unraveling the Causes of Deadly Epidemics
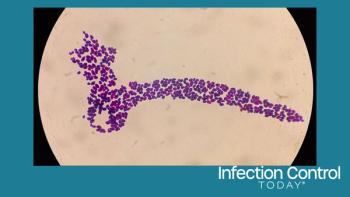
A team of collaborating scientists at The Methodist Hospital Research Institute in Houston, the Broad Institute in Boston, Mount Sinai Hospital in Toronto, and the Ontario Agency for Health Protection and Promotion (OAHPP) have sequenced almost 100 full genomes from three successive epidemics of flesh-eating bacteria. This has resulted in the first precise explanation of the biological events contributing to deadly epidemics of severe infection. This method can be used to track and help prevent devastating epidemics in the future.
Results of this research, funded by the National Institutes of Health and The Methodist Hospital Research Institute, appear in a study published online this week in Proceedings of the National Academy of Sciences (PNAS).
“The extensive full-genome data provide us with new clues about the bacteria’s ability to take advantage of vulnerabilities in the person who has contracted the bacteria,” said Dr. James M. Musser, co-director of The Methodist Hospital Research Institute and the senior corresponding author of the study. “With this type of unique molecular portrait of the bacterial pathogen, we can more effectively develop drugs to prevent the spread of epidemics and construct novel diagnostic and treatment strategies.”
“Until now, it has been a mystery why sometimes we see two opposing types of infection in patients who appear to have the same strain of flesh-eating bacteria,” said Dr. Donald Low, chief microbiologist at Mount Sinai Hospital, medical director of the OAHPP Public Health Laboratories and one of the authors of the study. “In some cases, patients suffer from a devastating infection of tissue and muscle requiring extensive surgery. And, other patients present with a skin infection readily treated with antibiotics. Now, we understand in part why this happens.”
Understanding the detailed molecular architecture of bacterial epidemics has been a long-sought goal of infectious disease research, he added. Genetics and evolution research has long been hobbled by the lack of comprehensive genome-wide markers. The recent advent of massively parallel DNA sequencing techniques now permits full-genome sequences to be generated rapidly. This opens the door to answering longstanding but previously economically unviable questions in all areas of biomedical research, including bacterial epidemics.
Musser’s lab, working closely with collaborators at Mount Sinai Hospital, the Ontario Provincial Public Health Laboratory, The Broad Institute in Boston and Sequenom in San Diego used short-read-length DNA sequencing coupled with mass spectroscopy analysis of single nucleotide polymorphisms (SNPs), to study the molecular pathogenomics of three successive epidemics of invasive infections involving 344 serotype M3 group A Streptococcus in Ontario, Canada. Sequencing the genome of 95 strains from the three epidemics, coupled with analysis of 280 bi-allelic SNPs in all 344 strains, revealed an unexpectedly complex population structure composed of a dynamic mixture of distinct clonally-related complexes. On average, each strain was differentiated from one another by only 49 SNPs and 11 insertion-deletion events (indels) in the core genome. The investigators discovered that each strain has a unique genome sequence, which brings unparalleled ability to track strain spread. The extensive full-genome data permitted the investigators to identify genes with unusually high rates of genetic variation, thereby providing new clues about selective forces at work in the host.