Researchers Untangle Genetics of Drug-Resistant TB
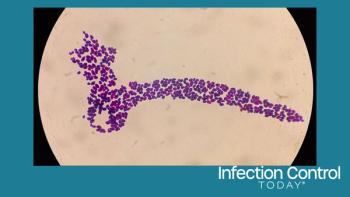
For years, physicians around the world have watched as strain after strain of the deadly bacteria Mycobacterium tuberculosis evolves resistance to drugs. Over the last few decades researchers have used the tools of molecular biology to identify a handful of individual mutations that allow TB to withstand many of the key therapeutics that doctors use to treat it. These genetic markers serve as clues for new drug development and as tools for diagnosing drug-resistant strains of TB. But the pace of discovery has proven too slow in the face of the complex array of rapidly mutating bacterial strains.
A new method of analyzing whole genome sequences of TB, applied to a massive set of strains of the bacteria collected from clinics around the world, has revealed 39 new genes associated with elevated drug resistance. The results were published Sept. 1, 2013 in Nature Genetics.
"We have found that more genes might be implicated in resistance than previously thought, and this means that we can start to unravel the role of these genes," says Megan Murray, HMS professor of global health and social medicine. "This is significant because it implicates new mechanisms in the evolution of resistance that can be further studied now and raises the possibility of more specific targets for the detection of resistance through molecular methods."
These new data suggest that acquiring resistance is a multistep process, perhaps requiring low-level resistance mutations prior to the ones that are well known. The findings also suggest that some of these new genes are involved in resistance that may confer "global" resistance traits, helping strains become resistant to a group of antibiotics rather than just one or a single class.
"Several of the genes we identified are related to the bacteria's regulation of cell walls; since many classes of drugs target the cell walls, we speculate that changes to the structure or metabolism of the cell walls might confer resistance to a wide variety of drugs," saya first author Maha Farhat, HMS instructor in medicine and assistant physician at Massachusetts General Hospital.
"Until now, people assumed that single mutations conferred high-level resistancea strain either had them or did notbut our results challenge that paradigm," Murray says. "Knowing that small changes early in the evolution of resistance open the door for big changes, or that a single change is a gateway to global resistance, would be important clues in our struggle to outrace evolving drug resistance."
Murray is part of a network of researchers and physicians working to develop a holistic, integrative approach to understanding and treating TB. In addition to her role as a researcher at HMS, she is also an associate professor of medicine at Brigham and Women's Hospital and professor in the Department of Epidemiology in the Harvard School of Public Health (HSPH), director of research at Partners in Health and director of the HMS Global Health and Social Medicine Research Core.
"We're not only implementing programs and documenting outcomes, we're using our access to clinics around the world to further basic scientific research, which will ultimately help improve standards of care," Murray says.
To find the novel drug-resistance genes, Farhat, Murray and collaborators adapted tools from evolutionary biology known as "phylogenetics." Phylogenetics allows the study of relationships within populations of organisms. It was originally developed to trace the path of evolution and to calculate how once-related organisms diverged onto different branches of the tree of life over many tens of thousands of years.
The team adapted these tools to measure the rapid-fire evolution of drug-resistant TB in the clinic.
They examined the whole genomes of 116 newly sequenced and 7 previously sequenced strains of TB. The sample included 47 strains with various levels of resistance to a variety of anti-TB drugs, as well as a group of susceptible strains, to allow the researchers to assess the genetic diversity of TB in the wild.
The method allows researchers to focus on the moments when resistant and susceptible strains branch off from one another. This silences the background noise of random mutations that aren't associated with resistance.
"You're tuning out all the changes that happened to their common ancestors, which allows you to look specifically at what the differences between the daughter lineages were when resistance evolved," Farhat says.
The project used a large set of clinical strains collected from human populations rather than strains that were developed in the lab. They sampled strains from outbreaks in British Columbia, Rome, South Africa and Russia. The strains came from dozens of sites around the world, including most of the major lineages of susceptible TB currently circulating around the world. This large, diverse data set was crucial to gaining insight into how resistant strains evolve in human populations.
They also needed a community of clinicians and researchers who were prepared to work across disciplines
"Making progress on understanding and fighting complex, global diseases like TB requires a community of physicians and scientists who are not only each working in their own niches, but building a collaborative ecosystem to share data, perspectives and results in order to push the work forward," Murray says.
One key collaboration was with Eric Rubin, HSPH professor of immunology and infectious diseases, and Karen Keiser, a graduate research fellow at HSPH. One of the new sites of resistance was found in ponA1, a gene that is important to TB cell wall function. Rubin and Keiser introduced this mutation into laboratory TB strains and found a small yet significant elevation in their levels of resistance to the TB drug rifampicin.
The small elevation in the resistance levels suggest that resistance is more complex than previously recognized and that likely multiple small mutations may act synergistically to result in full-blown resistance.
Drug resistance is a huge problem in TB, Murray says. In some settings in Eastern Europe, up to half of TB is multi-drug resistant. Extensively resistant strains have evolved that are very difficult to treat, and some strains in India and Iraq are virtually untreatable.
"We don't really understand why resistance develops so consistently," Murray says. "This study may provide a lens that we can use to see a way to develop better diagnostics for impending resistance, or even ways to prevent it from happening."
This study was funded by the Senior Ellison Foundation and the Massachusetts General Hospital Division of Pulmonary and Critical Care.
Source: Harvard Medical School Â